GRIN2C Antibodies
Background
The GRIN2C gene encodes a protein named glutamate ionotropic receptor NMDA type subunit 2C. This protein is mainly distributed in the granule cell layer of the central nervous system, especially in the cerebellum. This subunit, as a component of the NMDA receptor, plays a crucial role in learning, memory, and neural development by regulating glutamate-mediated synaptic transmission and synaptic plasticity. Abnormal function of the NMDA receptor is closely related to various neurological diseases. Therefore, the GRIN2C gene has attracted much attention in the research of neuro-psychiatric diseases. The cloning and functional analysis of this gene are facilitated by the advancement of molecular biology techniques, especially the in-depth study of its structure and regulatory mechanisms. This has greatly enhanced our understanding of glutamate-activated synaptic transmission, receptor heterogeneity, and their integration mechanisms in neural networks, providing important theoretical basis for the development of therapeutic targets for neurological diseases.
Structure of GRIN2C
The protein encoded by the GRIN2C gene has a molecular weight of approximately 13.7 kDa. There is a slight variation among different species due to sequence differences. The following table lists the parameters of this protein in several model organisms:
| Species | Human | Mouse | Rat | Chicken |
| Molecular Weight (kDa) | 13.7 | 13.8 | 13.8 | 13.6 |
| Primary Structural Differences | Containing four transmembrane domains, it constitutes an ion channel | The homology with human is up to 98% | The C-terminal domain is involved in synaptic localization | Displays differential expression during development |
The GluN2C subunit encoded by GRIN2C contains approximately 124 amino acids and mainly forms an extracellular N-terminal domain, four transmembrane segments, and an intracellular C-terminal tail. This protein assembles with other NMDA receptor subunits to form a hetero-tetrameric ion channel. The transmembrane regions form a selective pore that allows calcium ion influx. The extracellular domain contains a glycine binding site, while specific residues in the transmembrane region determine the gating properties and magnesium ion sensitivity of the channel. There are multiple phosphorylation sites in the protein structure, which regulate the channel activity and affect the synaptic transmission efficiency.
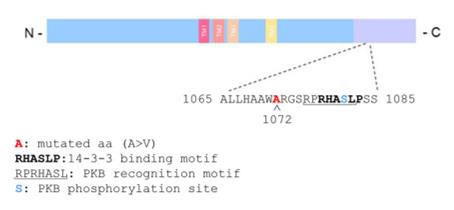 Fig. 1 Schematic representation of the GluN2C protein structure.1
Fig. 1 Schematic representation of the GluN2C protein structure.1
Key structural properties of GRIN2C:
- Encodes the GluN2C subunit, containing four transmembrane domains
- The extracellular N-terminal domain forms a ligand binding site
- The transmembrane regions constitute the ion channel pore
- The intracellular C-terminal contains a phosphorylation regulatory site
- Assembly with other NMDA receptor subunits into heterotetramers that mediate calcium influx
Functions of GRIN2C
The GluN2C subunit encoded by the GRIN2C gene mainly regulates the channel properties and synaptic functions of NMDA receptors, and is involved in various neurophysiological processes:
| Function | Description |
| Synaptic Transmission Regulation | As a component subunit of the NMDA receptor, it mediates calcium ion influx, triggering downstream signal cascades. |
| Receptor Kinetic Regulation | Endows the NMDA receptor with unique gating characteristics, including slower channel inactivation and low magnesium ion sensitivity. |
| Contribution to Synaptic Plasticity | Participates in the induction of long-term potentiation and long-term depression, influencing the learning and memory process. |
| Neurodevelopmental Support | Expresses in specific brain regions during development, regulating neuronal migration and synaptic formation. |
| Neuroprotection and Injury | Moderate activation promotes the survival of neurons, while excessive activation leads to excitotoxic damage. |
The activation of NMDA receptors requires the simultaneous binding of glutamate and glycine. The presence of the GluN2C subunit makes the receptor less sensitive to glutamate but more sensitive to glycine. This unique pharmacological property enables it to play a special role in the regulation of synaptic transmission.
Applications of GRIN2C and GRIN2C Antibody in Literature
1. Rubino, Elisa, et al. "Exome sequencing reveals a rare damaging variant in GRIN2C in familial late-onset Alzheimer's disease." Alzheimer's research & therapy 17.1 (2025): 21. https://doi.org/10.1186/s13195-024-01661-y
In this study, a rare missense variant p.(A1072V) in the GRIN2C gene was identified in an Italian late-onset Alzheimer's disease family. This variant enhances NMDAR current and membrane expression, and is co-localized with 14-3-3 protein, suggesting that abnormal glutamatergic signaling may be involved in the disease mechanism.
2. Xia, Yu, et al. "Inflammatory factor IL1α induces aberrant astrocyte proliferation in spinal cord injury through the Grin2c/Ca2+/CaMK2b pathway." Neuroscience Bulletin 40.4 (2024): 421-438. https://doi.org/10.1007/s12264-023-01128-4
The study found that after spinal cord injury, IL1α inhibits the Grin2c/Ca2+/CaMK2b pathway, thereby promoting abnormal proliferation of astrocytes. Neutralizing IL1α can upregulate Grin2c and inhibit this process, providing a new target for spinal cord injury repair.
3. Hernandez, Giovanni, et al. "Dorsal raphe stimulation relays a reward signal to the ventral tegmental area via GluN2C NMDA receptors." Plos one 18.11 (2023): e0293564. https://doi.org/10.1371/journal.pone.0293564
The study found that the ventral tegmental area (VTA) of the midbrain is rich in the GluN2C subunit, and approximately half of the dopamine neurons express its gene. Downregulation of this subunit weakens the brain's stimulus reward effect, revealing its crucial role in mediating the glutamatergic reward signal transmission from the nucleus accumbens to the VTA.
4. Shelkar, Gajanan P., et al. "Differential effect of NMDA receptor GluN2C and GluN2D subunit ablation on behavior and channel blocker-induced schizophrenia phenotypes." Scientific Reports 9.1 (2019): 7572. https://doi.org/10.1038/s41598-019-43957-2
This study utilized pure knockout mice to compare the functions of the GluN2C/D subunits. The GluN2C knockout mice were insensitive to PCP-induced hyperactivity, and the effect of GluN2C/D enhancers in improving PPI defects was dependent on GluN2C, revealing its unique role in schizophrenia-like behaviors.
5. Teng, Huajing, et al. "Evolutionary mode and functional divergence of vertebrate NMDA receptor subunit 2 genes." PloS one 5.10 (2010): e13342. https://doi.org/10.1371/journal.pone.0013342
The research reveals that the GRIN2 gene family in vertebrates generates four subtypes through two rounds of genome duplication, while fish have a unique duplication that produces additional copies. The functional divergence mainly occurs after the first round of duplication, with purifying selection playing a dominant role in its evolution, providing new insights into the mechanism of cognitive evolution.
Creative Biolabs: GRIN2C Antibodies for Research
Creative Biolabs specializes in the production of high-quality GRIN2C antibodies for research and industrial applications. Our portfolio includes monoclonal and polyclonal antibodies tailored for ELISA, Flow Cytometry, Western blot, immunohistochemistry, and other diagnostic methodologies.
- Custom GRIN2C Antibody Development: Tailor-made solutions to meet specific research requirements.
- Bulk Production: Large-scale antibody manufacturing for industry partners.
- Technical Support: Expert consultation for protocol optimization and troubleshooting.
- Aliquoting Services: Conveniently sized aliquots for long-term storage and consistent experimental outcomes.
For more details on our GRIN2C antibodies, custom preparations, or technical support, contact us at email.
Reference
- Rubino, Elisa, et al. "Exome sequencing reveals a rare damaging variant in GRIN2C in familial late-onset Alzheimer's disease." Alzheimer's research & therapy 17.1 (2025): 21. Distributed under Open Access license CC BY 4.0, Cropped from the original figure. https://doi.org/10.1186/s13195-024-01661-y
Anti-GRIN2C antibodies
 Loading...
Loading...
Hot products 
-
Mouse Anti-CCN2 Recombinant Antibody (CBFYC-2383) (CBMAB-C2456-FY)

-
Mouse Anti-ARSA Recombinant Antibody (CBYC-A799) (CBMAB-A3679-YC)

-
Mouse Anti-ADAM29 Recombinant Antibody (V2-179787) (CBMAB-A1149-YC)

-
Mouse Anti-AAV-5 Recombinant Antibody (V2-503417) (CBMAB-V208-1369-FY)

-
Mouse Anti-COL12A1 Recombinant Antibody (CBYY-C3117) (CBMAB-C4560-YY)

-
Mouse Anti-CCND2 Recombinant Antibody (DCS-3) (CBMAB-G1318-LY)

-
Rabbit Anti-CCL5 Recombinant Antibody (R0437) (CBMAB-R0437-CN)

-
Mouse Anti-AAV-5 Recombinant Antibody (V2-503416) (CBMAB-V208-1402-FY)

-
Rat Anti-FABP3 Recombinant Antibody (CBXF-2299) (CBMAB-F1612-CQ)

-
Mouse Anti-AMIGO2 Recombinant Antibody (CBYY-C0756) (CBMAB-C2192-YY)

-
Mouse Anti-ADIPOR2 Recombinant Antibody (V2-179983) (CBMAB-A1369-YC)

-
Mouse Anti-CD83 Recombinant Antibody (HB15) (CBMAB-C1765-CQ)

-
Mouse Anti-ACTN4 Recombinant Antibody (V2-6075) (CBMAB-0020CQ)

-
Mouse Anti-AZGP1 Recombinant Antibody (CBWJZ-007) (CBMAB-Z0012-WJ)

-
Mouse Anti-GIPC2 Recombinant Antibody (10) (CBMAB-G0476-LY)

-
Mouse Anti-AMH Recombinant Antibody (5/6) (CBMAB-A2527-YC)

-
Mouse Anti-GFAP Recombinant Antibody (24) (CBMAB-G2927-LY)

-
Rabbit Anti-ALOX5AP Recombinant Antibody (CBXF-1219) (CBMAB-F0750-CQ)

-
Mouse Anti-ARHGDIA Recombinant Antibody (CBCNA-009) (CBMAB-R0415-CN)

-
Mouse Anti-CCNH Recombinant Antibody (CBFYC-1054) (CBMAB-C1111-FY)

- AActivation
- AGAgonist
- APApoptosis
- BBlocking
- BABioassay
- BIBioimaging
- CImmunohistochemistry-Frozen Sections
- CIChromatin Immunoprecipitation
- CTCytotoxicity
- CSCostimulation
- DDepletion
- DBDot Blot
- EELISA
- ECELISA(Cap)
- EDELISA(Det)
- ESELISpot
- EMElectron Microscopy
- FFlow Cytometry
- FNFunction Assay
- GSGel Supershift
- IInhibition
- IAEnzyme Immunoassay
- ICImmunocytochemistry
- IDImmunodiffusion
- IEImmunoelectrophoresis
- IFImmunofluorescence
- IGImmunochromatography
- IHImmunohistochemistry
- IMImmunomicroscopy
- IOImmunoassay
- IPImmunoprecipitation
- ISIntracellular Staining for Flow Cytometry
- LALuminex Assay
- LFLateral Flow Immunoassay
- MMicroarray
- MCMass Cytometry/CyTOF
- MDMeDIP
- MSElectrophoretic Mobility Shift Assay
- NNeutralization
- PImmunohistologyp-Paraffin Sections
- PAPeptide Array
- PEPeptide ELISA
- PLProximity Ligation Assay
- RRadioimmunoassay
- SStimulation
- SESandwich ELISA
- SHIn situ hybridization
- TCTissue Culture
- WBWestern Blot






