P4HB Antibodies
Background
The P4HB gene encodes a multifunctional endoplasmic reticulum-resident protein, which possesses both proline-4-hydroxylase activity and molecular chaperone function. Its main functions include catalyzing the proline hydroxylation reaction during collagen synthesis, as well as assisting in the correct folding and assembly of newly synthesized peptide chains, thereby maintaining the stability and function of the extracellular matrix. This gene plays a crucial role in various physiological processes, especially showing significant expression enhancement in tissue repair, embryonic development, and stress response scenarios. As a member of the protein disulfide isomerase family, it can also participate in redox homeostasis regulation. Its dysfunction is closely related to various fibrotic diseases, tumor progression, and metabolic abnormalities. Since its function was systematically elucidated, P4HB has become an important model molecule in cell biology and clinical medicine research, providing a key theoretical basis for understanding the endoplasmic reticulum quality control mechanism and the dynamic balance of the extracellular matrix.
Structure of P4HB
The protein encoded by the P4HB gene has a molecular weight of approximately 57 kDa. This value is highly conserved among different mammalian species, mainly due to the high consistency of the core functional domain sequence.
| Species | Human | Mouse | Rat |
| Molecular Weight (kDa) | 57.1 | 57.0 | 57.2 |
| Primary Structural Differences | Containing a signal peptide, a catalytic domain and a KEDL endoplasmic reticulum retention sequence | Very high homology with humans, the catalytic site completely conservative | The structures are highly similar and the functions can be interchangeable. |
This protein is composed of approximately 500 amino acid residues and its three-dimensional structure exhibits a typical multi-domain folding pattern. Its core function relies on two key active sites: one is the active center of proline 4-hydroxylase, which is responsible for catalyzing the hydroxylation reaction of proline in collagen; the other is the thioredoxin-like domain, which has the activity of protein disulfide isomerase and participates in the correct folding of peptide chains. These two active sites work in synergy spatially, enabling them to simultaneously participate in collagen synthesis and quality control. The KDEL sequence at the C-terminus of the protein is the key signal for its localization in the endoplasmic reticulum cavity.
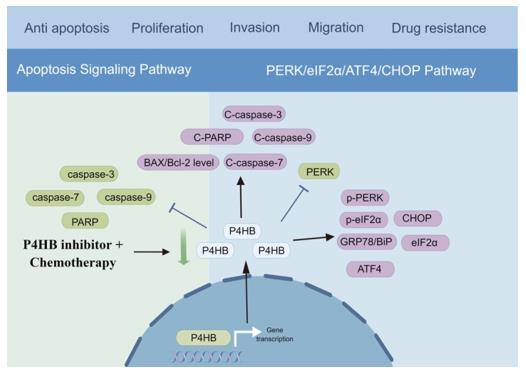 Fig. 1 The underlying mechanism of P4HB in bladder cancer.1
Fig. 1 The underlying mechanism of P4HB in bladder cancer.1
Key structural properties of P4HB:
- Multi-domain folded configuration
- Hydrophobic pockets form around the active site
- TRX sample structure domain contains CXXC motif
- The C-terminal KDEL ER retention sequence ensures its accurate localization to the ER lumen
Functions of P4HB
The core function of the P4HB protein is to act as a prolyl-4-hydroxylase to catalyze collagen synthesis, and it also has molecular chaperone activity to assist protein folding. Additionally, it is involved in various cellular processes, including endoplasmic reticulum stress response and regulation of redox homeostasis.
| Function | Description |
| Collagen Synthesis Catalysis | Catalyzes the hydroxylation reaction of proline residues in pre-collagen proteins, which is a crucial step in forming stable triple-helix collagen molecules. |
| Molecular Chaperone | By its disulfide bond isomerase activity, it assists in the correct folding and assembly of newly synthesized peptide chains, and prevents the formation of incorrect aggregates. |
| Endoplasmic Reticulum Quality Control | As an endoplasmic reticulum stress sensor, it participates in the unfolded protein response and maintains the protein homeostasis of organelles. |
| Redox Regulation | By means of its thioredoxin-like domain, it catalyzes the formation, reduction and isomerization of disulfide bonds, thereby regulating the intracellular redox state. |
| Cell Adhesion and Migration | By influencing the maturation and stability of the extracellular matrix, it indirectly regulates the adhesion, spreading and migration processes of cells. |
Unlike monofunctional enzymes, P4HB achieves a tight coupling of collagen biosynthesis and quality control processes in both space and time through the synergistic action of its dual-function active center, ensuring the efficient and highly accurate generation of extracellular matrix.
Applications of P4HB and P4HB Antibody in Literature
1. Ouyang, Fan, et al. "Aloe Emodin Alleviates Radiation‐Induced Heart Disease via Blocking P4HB Lactylation and Mitigating Kynurenine Metabolic Disruption." Advanced Science 11.47 (2024): 2406026. https://doi.org/10.1002/advs.202406026
This study reveals that aloin stabilizes mitochondrial GOT2 activity by inhibiting lactation at the K311 site of P4HB protein, thereby regulating the PTGS2/SH3GLB1/NDP52 axis to alleviate radiation-induced heart injury, providing a new strategy for targeted lactation in the prevention and treatment of heart injury.
2. Feng, Dechao, et al. "Targeting prolyl 4-hydroxylase subunit beta (P4HB) in cancer: new roads to travel." Aging and Disease 15.6 (2023): 2369. https://doi.org/10.14336/AD.2023.1126
This study reveals that prolyl 4-hydroxylase subunit β (P4HB) plays a promoting role in various cancers. This article reveals its carcinogenic mechanism through pan-cancer analysis, predicts related interacting molecules and potential targeted drugs, and provides new targets for tumor treatment.
3. Feng, Dechao, et al. "Prolyl 4-hydroxylase subunit beta (P4HB) could serve as a prognostic and radiosensitivity biomarker for prostate cancer patients." European Journal of Medical Research 28.1 (2023): 245. https://doi.org/10.1186/s40001-023-01215-2
This study reveals that prolyl 4-hydroxylase subunit β (P4HB) is significantly highly expressed in prostate cancer and is closely related to poor prognosis and radiotherapy resistance. Down-regulation of P4HB can effectively inhibit the proliferation of cancer cells, suggesting that it can serve as a potential prognostic marker and therapeutic target.
4. Dawoud, Marwa Mohammed, et al. "The Role of P4HB and SOX4 in Prostatic Carcinoma and Their Clinical Significance." Iranian Journal of Pathology 19.2 (2024): 225. https://doi.org/10.30699/IJP.2024.2017851.3227
This study confirmed that prolyl 4-hydroxylase subunit β (P4HB) and SOX4 are significantly upregulated in prostate cancer, and the two are positively correlated. They are closely related to tumor grade and invasiveness and can be used as potential diagnostic and prognostic markers for prostate cancer.
5. Tang, Yulian, et al. "Expression, localization, and function of P4HB in the spermatogenesis of Chinese mitten crab (Eriocheir sinensis)." PeerJ 11 (2023): e15547. https://doi.org/10.7717/peerj.15547
This study was the first to locate and analyze the P4HB protein during spermatogenesis in the Chinese mitten crab. The results show that the expression and localization of P4HB vary in male germ cells at different developmental stages, and it may play a key role in maintaining the stability of the non-condensable nucleus of sperm.
Creative Biolabs: P4HB Antibodies for Research
Creative Biolabs specializes in the production of high-quality P4HB antibodies for research and industrial applications. Our portfolio includes monoclonal antibodies tailored for ELISA, Flow Cytometry, Western blot, immunohistochemistry, and other diagnostic methodologies.
- Custom P4HB Antibody Development: Tailor-made solutions to meet specific research requirements.
- Bulk Production: Large-scale antibody manufacturing for industry partners.
- Technical Support: Expert consultation for protocol optimization and troubleshooting.
- Aliquoting Services: Conveniently sized aliquots for long-term storage and consistent experimental outcomes.
For more details on our P4HB antibodies, custom preparations, or technical support, contact us at email.
Reference
- Feng, Dechao, et al. "Targeting prolyl 4-hydroxylase subunit beta (P4HB) in cancer: new roads to travel." Aging and Disease 15.6 (2023): 2369. https://doi.org/10.14336/AD.2023.1126
Anti-P4HB antibodies
Products List
 Loading...
Loading...
Hot products 
-
Mouse Anti-CEMIP Recombinant Antibody (3C12) (CBMAB-K0296-LY)

-
Mouse Anti-4-Hydroxynonenal Recombinant Antibody (V2-502280) (CBMAB-C1055-CN)

-
Mouse Anti-CCS Recombinant Antibody (CBFYC-1093) (CBMAB-C1150-FY)

-
Mouse Anti-BZLF1 Recombinant Antibody (BZ.1) (CBMAB-AP705LY)

-
Mouse Anti-BLK Recombinant Antibody (CBYY-0618) (CBMAB-0621-YY)

-
Mouse Anti-DDC Recombinant Antibody (8E8) (CBMAB-0992-YC)

-
Mouse Anti-ACE2 Recombinant Antibody (V2-179293) (CBMAB-A0566-YC)

-
Mouse Anti-APCS Recombinant Antibody (CBYC-A663) (CBMAB-A3054-YC)

-
Human Anti-SARS-CoV-2 S1 Monoclonal Antibody (CBFYR-0120) (CBMAB-R0120-FY)

-
Rat Anti-CD63 Recombinant Antibody (7G4.2E8) (CBMAB-C8725-LY)

-
Mouse Anti-ARSA Recombinant Antibody (CBYC-A799) (CBMAB-A3679-YC)

-
Rabbit Anti-B2M Recombinant Antibody (CBYY-0059) (CBMAB-0059-YY)

-
Mouse Anti-FOXA3 Recombinant Antibody (2A9) (CBMAB-0377-YC)

-
Rabbit Anti-ENO2 Recombinant Antibody (BA0013) (CBMAB-0272CQ)

-
Mouse Anti-AMIGO2 Recombinant Antibody (CBYY-C0756) (CBMAB-C2192-YY)

-
Mouse Anti-CIITA Recombinant Antibody (CBLC160-LY) (CBMAB-C10987-LY)

-
Rabbit Anti-ALOX5AP Recombinant Antibody (CBXF-1219) (CBMAB-F0750-CQ)

-
Mouse Anti-ASH1L Monoclonal Antibody (ASH5H03) (CBMAB-1372-YC)

-
Mouse Anti-CCDC6 Recombinant Antibody (CBXC-0106) (CBMAB-C5397-CQ)

-
Mouse Anti-G6PD Recombinant Antibody (13B331) (CBMAB-G1553-LY)

- AActivation
- AGAgonist
- APApoptosis
- BBlocking
- BABioassay
- BIBioimaging
- CImmunohistochemistry-Frozen Sections
- CIChromatin Immunoprecipitation
- CTCytotoxicity
- CSCostimulation
- DDepletion
- DBDot Blot
- EELISA
- ECELISA(Cap)
- EDELISA(Det)
- ESELISpot
- EMElectron Microscopy
- FFlow Cytometry
- FNFunction Assay
- GSGel Supershift
- IInhibition
- IAEnzyme Immunoassay
- ICImmunocytochemistry
- IDImmunodiffusion
- IEImmunoelectrophoresis
- IFImmunofluorescence
- IGImmunochromatography
- IHImmunohistochemistry
- IMImmunomicroscopy
- IOImmunoassay
- IPImmunoprecipitation
- ISIntracellular Staining for Flow Cytometry
- LALuminex Assay
- LFLateral Flow Immunoassay
- MMicroarray
- MCMass Cytometry/CyTOF
- MDMeDIP
- MSElectrophoretic Mobility Shift Assay
- NNeutralization
- PImmunohistologyp-Paraffin Sections
- PAPeptide Array
- PEPeptide ELISA
- PLProximity Ligation Assay
- RRadioimmunoassay
- SStimulation
- SESandwich ELISA
- SHIn situ hybridization
- TCTissue Culture
- WBWestern Blot






