BET1 Antibodies
Background
The protein encoded by the BET1 gene is a key lipid transporter located on the cell membrane and endoplasmic reticulum. It plays a central role in the endoplasmic reticulum-Golgi transport system of mammalian cells. As a member of the phospholipid transfer protein family, this protein can specifically bind and transport phospholipid molecules, maintaining the lipid composition balance of organelle membranes and ensuring the synthesis and secretion processes of proteins. The occurrence of diseases such as polycystic kidney disease is closely related to the abnormal function of the BET1 gene. When lipid transport is blocked, it will lead to the disorder of the intracellular material transport system. This gene was initially discovered in yeast genetic screening, and subsequent studies have confirmed its conserved function in the endoplasmic reticulum transport system of mammalian cells. After the unique lipid-binding domain and membrane localization mechanism were deeply analyzed, it provided an important model for revealing the molecular regulatory network of intracellular membrane system transport, greatly promoting our understanding of cell homeostasis maintenance and vesicle transport mechanisms.
Structure of BET1
The protein encoded by the BET1 gene has a molecular weight of approximately 22 kDa. There are slight differences among different species, and the specific data are shown in the table below:
| Species | Human | Mouse | Rat | Zebrafish |
| Molecular Weight (kDa) | 22.3 | 22.1 | 22.1 | 21.8 |
| Primary Structural Differences | Highly conservative | Homology: 98% | Homology 97% | Evolutionarily conserved |
This protein is composed of approximately 200 amino acids. Its three-dimensional structure contains a characteristic long intramolecular domain, which mediates protein interactions through a coiled-coil structure. The BET1 protein is located on the surface of the Golgi membrane. Its structure contains a conserved C-terminal transmembrane domain, which anchors the protein to the membrane of transport vesicles. The cytoplasmic region of the protein contains multiple coiled-coil motifs, which form specific hydrophobic regions and mediate the binding with other members of the SNARE complex. The key glutamine residues are located at the center of the SNARE motif, which stabilize the conformation of the fusion complex by forming an ionic layer, thereby precisely regulating the specific recognition and fusion process between vesicles and the target membrane.
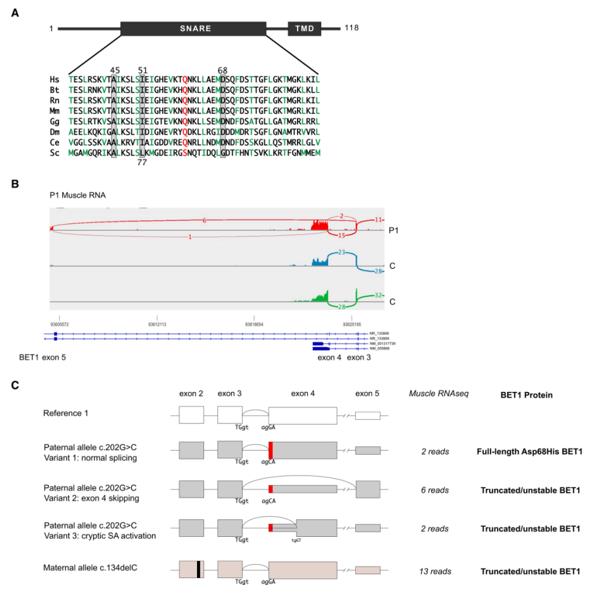 Fig. 1 RNA sequencing analysis of the BET1 c.202G>C; p.(Asp68His) missense variant.1
Fig. 1 RNA sequencing analysis of the BET1 c.202G>C; p.(Asp68His) missense variant.1
Key structural properties of BET1:
- The characteristic SNARE coiled-coil domain
- The hydrophobic interface mediates the assembly of protein complexes
- The glutamine residue stabilizes the fusion complex
- The coiled-coil motif in the cytoplasmic region regulates vesicle recognition
Functions of BET1
The main function of the BET1 gene is to mediate the vesicle transport between the endoplasmic reticulum and the Golgi apparatus, playing a central role in the intracellular material transport process. However, it is also involved in various cellular biological processes, including maintaining the structure of the Golgi apparatus and regulating autophagy.
| Function | Description |
| Vesicle transport | As a SNARE protein, BET1 mediates the recognition and fusion of vesicles derived from the endoplasmic reticulum with the Golgi membrane, ensuring the normal transport of proteins from the endoplasmic reticulum to the Golgi apparatus. |
| Golgi apparatus homeostasis | It participates in maintaining the integrity of the Golgi apparatus structure, influencing the morphology and function of the Golgi apparatus by regulating the balance of membrane vesicle transport. |
| Autophagy regulation | It assists in the formation of autophagosomes during autophagy and participates in the removal of cellular waste and nutrient cycling through membrane transport mechanisms. |
| Cell stress response | It regulates the membrane transport flux under stress conditions, helping cells adapt to endoplasmic reticulum stress and nutrient deficiency states. |
| Secretion pathway maintenance | It ensures the normal transport of secretory proteins and membrane proteins, maintaining the stability of the cell's secretion function. |
The membrane fusion process mediated by BET1 has strict temporal and spatial specificity. It ensures the accurate targeting of transport vesicles to the Golgi membrane by forming a trans-complex with other SNARE proteins, demonstrating its precise regulatory role in the directional transport of substances within the cell.
Applications of BET1 and BET1 Antibody in Literature
1. Donkervoort, Sandra, et al. "BET1 variants establish impaired vesicular transport as a cause for muscular dystrophy with epilepsy." EMBO Molecular Medicine 13.12 (2021): EMMM202013787. https://doi.org/10.15252/emmm.202013787
This study is the first to reveal that mutations in the BET1 gene cause congenital muscular dystrophy. The variations interfere with the BET1 protein and its interaction with ERGIC-53, causing endoplasmic reticulum-Golgi transport disorders, providing new insights into the disease mechanism.
2. Garcia, Gilberto, et al. "Large‐scale genetic screens identify BET‐1 as a cytoskeleton regulator promoting actin function and life span." Aging Cell 22.1 (2023): e13742. https://doi.org/10.1111/acel.13742
The study found that BET-1 regulates the protection of the actin cytoskeleton through transcription, and delays the functional decline associated with aging. Overexpression of bet-1 in nematodes can maintain the actin function in old age, significantly prolong lifespan and health span, revealing a new mechanism for cytoskeleton homeostasis.
3. Gee, Fiona, et al. "An RNAi-based dimorphic genetic screen identified the double bromodomain protein BET-1 as a sumo-dependent attenuator of RAS-mediated signalling." Plos one 8.12 (2013): e83659. https://doi.org/10.1371/journal.pone.0083659
The study found that BET-1 acts as a substrate for SUMOylation modification and regulates gene expression by physically binding to factors such as SMO-1, thereby inhibiting excessive activation of the RAS/MAPK signaling pathway. This mechanism reveals a new mode of the collaborative interaction between chromatin regulation and SUMOylation.
4. Zhang, Tao, and Wanjin Hong. "Ykt6 forms a SNARE complex with syntaxin 5, GS28, and Bet1 and participates in a late stage in endoplasmic reticulum-Golgi transport." Journal of Biological Chemistry 276.29 (2001): 27480-27487. https://doi.org/10.1074/jbc.M102786200
The research has confirmed that Ykt6 is located in the anterograde Golgi apparatus and forms a SNARE complex with proteins such as Bet1, mediating the late fusion of ER-Golgi transport. The functional blockade of Ykt6 leads to fragmentation of the Golgi apparatus, revealing the unique two-stage sequential mechanism of SNARE complex in mammalian cells.
5. Lawal, Bashir, et al. "Transcriptomic-based identification of the immuno-oncogenic signature of cholangiocarcinoma for HLC-018 multi-target therapy exploration." Cells 10.11 (2021): 2873. https://doi.org/10.3390/cells10112873
The research has found that five key genes such as BET1 are highly expressed in cholangiocarcinoma, and they are closely related to tumor immune suppression, metastasis and drug resistance. The small molecule HLC-018 can efficiently bind to these targets, providing a new strategy for the treatment of cholangiocarcinoma.
Creative Biolabs: BET1 Antibodies for Research
Creative Biolabs specializes in the production of high-quality BET1 antibodies for research and industrial applications. Our portfolio includes monoclonal and polyclonal antibodies tailored for ELISA, Flow Cytometry, Western blot, immunohistochemistry, and other diagnostic methodologies.
- Custom BET1 Antibody Development: Tailor-made solutions to meet specific research requirements.
- Bulk Production: Large-scale antibody manufacturing for industry partners.
- Technical Support: Expert consultation for protocol optimization and troubleshooting.
- Aliquoting Services: Conveniently sized aliquots for long-term storage and consistent experimental outcomes.
For more details on our BET1 antibodies, custom preparations, or technical support, contact us at email.
Reference
- Donkervoort, Sandra, et al. "BET1 variants establish impaired vesicular transport as a cause for muscular dystrophy with epilepsy." EMBO Molecular Medicine 13.12 (2021): EMMM202013787. Distributed under Open Access license CC BY 4.0, without modification. https://doi.org/10.15252/emmm.202013787
Anti-BET1 antibodies
 Loading...
Loading...
Hot products 
-
Mouse Anti-DES Monoclonal Antibody (440) (CBMAB-AP1857LY)

-
Mouse Anti-CD24 Recombinant Antibody (2Q1282) (CBMAB-C1624-CN)

-
Mouse Anti-CD24 Recombinant Antibody (SN3) (CBMAB-C1037-CQ)

-
Mouse Anti-ATM Recombinant Antibody (2C1) (CBMAB-A3970-YC)

-
Rat Anti-(1-5)-α-L-Arabinan Recombinant Antibody (V2-501861) (CBMAB-XB0003-YC)

-
Mouse Anti-G6PD Recombinant Antibody (13B331) (CBMAB-G1553-LY)

-
Mouse Anti-CASP7 Recombinant Antibody (10-01-62) (CBMAB-C2005-LY)

-
Rabbit Anti-ADRA1A Recombinant Antibody (V2-12532) (CBMAB-1022-CN)

-
Rabbit Anti-DLK1 Recombinant Antibody (9D8) (CBMAB-D1061-YC)

-
Mouse Anti-C5b-9 Recombinant Antibody (aE11) (CBMAB-AO138LY)

-
Rabbit Anti-BAD (Phospho-Ser136) Recombinant Antibody (CAP219) (CBMAB-AP536LY)

-
Mouse Anti-CD24 Recombinant Antibody (HIS50) (CBMAB-C10123-LY)

-
Mouse Anti-AKT1 Recombinant Antibody (V2-180546) (CBMAB-A2070-YC)

-
Mouse Anti-CAT Recombinant Antibody (724810) (CBMAB-C8431-LY)

-
Mouse Anti-AK4 Recombinant Antibody (V2-180419) (CBMAB-A1891-YC)

-
Mouse Anti-CCNH Recombinant Antibody (CBFYC-1054) (CBMAB-C1111-FY)

-
Mouse Anti-FAS2 Monoclonal Antibody (1D4) (CBMAB-0071-CN)

-
Mouse Anti-AAV8 Recombinant Antibody (V2-634028) (CBMAB-AP022LY)

-
Mouse Anti-DLL4 Recombinant Antibody (D1090) (CBMAB-D1090-YC)

-
Mouse Anti-ARHGAP5 Recombinant Antibody (54/P190-B) (CBMAB-P0070-YC)

- AActivation
- AGAgonist
- APApoptosis
- BBlocking
- BABioassay
- BIBioimaging
- CImmunohistochemistry-Frozen Sections
- CIChromatin Immunoprecipitation
- CTCytotoxicity
- CSCostimulation
- DDepletion
- DBDot Blot
- EELISA
- ECELISA(Cap)
- EDELISA(Det)
- ESELISpot
- EMElectron Microscopy
- FFlow Cytometry
- FNFunction Assay
- GSGel Supershift
- IInhibition
- IAEnzyme Immunoassay
- ICImmunocytochemistry
- IDImmunodiffusion
- IEImmunoelectrophoresis
- IFImmunofluorescence
- IGImmunochromatography
- IHImmunohistochemistry
- IMImmunomicroscopy
- IOImmunoassay
- IPImmunoprecipitation
- ISIntracellular Staining for Flow Cytometry
- LALuminex Assay
- LFLateral Flow Immunoassay
- MMicroarray
- MCMass Cytometry/CyTOF
- MDMeDIP
- MSElectrophoretic Mobility Shift Assay
- NNeutralization
- PImmunohistologyp-Paraffin Sections
- PAPeptide Array
- PEPeptide ELISA
- PLProximity Ligation Assay
- RRadioimmunoassay
- SStimulation
- SESandwich ELISA
- SHIn situ hybridization
- TCTissue Culture
- WBWestern Blot






