IDH2 Antibodies
Background
The IDH2 gene encodes a mitochondrial protein called isocitrate dehydrogenase 2. This enzyme is mainly involved in the tricarboxylic acid cycle and catalyzes the oxidation decarboxylation of isocitrate to form α-ketoglutarate. In various tumors (such as gliomas and leukemia), IDH2 often undergoes specific site mutations, resulting in abnormal enzyme function and the accumulation of carcinogenic metabolite 2-hydroxyglutaric acid, thereby affecting cellular epigenetic modification and differentiation processes. Since its tumor-related mutations were discovered in 2009, IDH2 has become an important target in cancer metabolism research. Inhibitors targeting its mutant forms have been applied in the clinical treatment of some hematological tumors. The systematic study of this gene has deepened our understanding of metabolic reprogramming and the mechanism of tumor occurrence, and has promoted the development of targeted metabolic therapies.
Structure of IDH2
The IDH2 gene encodes the isocitrate dehydrogenase 2 protein, which has a molecular weight of approximately 47 kDa. Its size is relatively conserved among different species, mainly due to the highly stable catalytic core structure.
| Species | Human | Mouse | Rat |
| Molecular Weight (kDa) | ~47 | ~47 | ~47 |
| Primary Structural Differences | Common cancer-causing mutations exist at sites such as R132 and R172. | Highly similar to humans and often used for constructing disease models | Highly conserved structure, used for basal metabolism research |
This protein is composed of 452 amino acids and forms a homodimeric structure. Its tertiary structure consists of three key domains: the substrate binding domain, the coenzyme NADP⁺binding domain, and the dimer interface domain. The active center is located at the junction of the domains, and the key arginine residues (such as R172) directly participate in the catalysis. Its secondary structure is mainly composed of α helices and β sheets, which together form a hydrophobic channel to stabilize the binding of isocitrate substrate and NADP⁺ coenzyme.
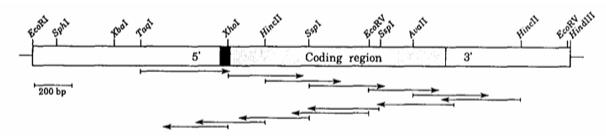 Fig. 1 Restriction map and sequencing strategy for the IDH2 gene.1
Fig. 1 Restriction map and sequencing strategy for the IDH2 gene.1
Key structural properties of IDH2:
- Forming the folded structure of homologous dimers
- The active catalytic center is composed of three domains.
- NADP⁺-dependent coenzyme and divalent metal ion binding sites
Functions of IDH2
The core function of the protein encoded by the IDH2 gene is to catalyze the oxidation decarboxylation of isocitrate to produce α-ketoglutarate (α-KG). However, this protein can also cause abnormal accumulation of carcinogenic metabolites under specific mutant conditions.
| Function | Description |
| Metabolic Catalysis | Catalyzes the conversion of isocitrate to α-ketoglutarate in the mitochondrial tricarboxylic acid cycle, and reduces NADP⁺to NADPH. |
| Metabolic Reprogramming | After mutations at sites such as R172 or R132, new functions are acquired, enabling the reduction of α-KG to the carcinogenic metabolite 2-hydroxyglutaric acid (2-HG). |
| Epigenetic Regulation | The abnormal accumulation of 2-HG competitively inhibits α-KG-dependent dioxygenases (such as TET and histone demethylases), interfering with cellular epigenetic modifications. |
| Dysdifferentiation Block | Especially in hematopoietic cells, the mutation leads to the accumulation of 2-HG, which hinders the normal differentiation process of the cells and promotes the occurrence of leukemia. |
| Redox Equilibrium | The wild-type IDH2 helps maintain the intracellular redox balance and resist oxidative stress by generating NADPH. |
The enzymatic kinetic characteristics of the mutant IDH2 undergo fundamental changes, with its catalytic reaction shifting from oxidative decarboxylation to reductive carboxylation. This "functional gain" mutation is a metabolic driving factor for the occurrence of various tumors.
Applications of IDH2 and IDH2 Antibody in Literature
1. Cupp, J. R., and L. McAlister-Henn. "NAD (+)-dependent isocitrate dehydrogenase. Cloning, nucleotide sequence, and disruption of the IDH2 gene from Saccharomyces cerevisiae." Journal of Biological Chemistry 266.33 (1991): 22199-22205. https://doi.org/10.1016/S0021-9258(18)54554-3
The article indicates that the NAD+ dependent isocitrate dehydrogenase of Saccharomyces cerevisiae consists of the IDH1 and IDH2 subunits. The IDH2 gene encodes a mitochondrial-targeted precursor, which, together with IDH1, is necessary for the enzyme's activity. The absence of this enzyme prevents cells from utilizing acetic acid and affects the function of the tricarboxylic acid cycle.
2. Issa, Ghayas C., and Courtney D. DiNardo. "Acute myeloid leukemia with IDH1 and IDH2 mutations: 2021 treatment algorithm." Blood cancer journal 11.6 (2021): 107. https://doi.org/10.1038/s41408-021-00497-1
The article indicates that approximately 20% of acute myeloid leukemia cases carry mutations in the IDH1/2 genes, which lead to the production of the carcinogenic metabolite 2-hydroxyglutarate and block myeloid differentiation. Selective oral IDH inhibitors have been approved for monotherapy and are currently being evaluated in combination with chemotherapy and targeted therapies to enhance efficacy.
3. Zou, Peng, et al. "IDH1/IDH2 mutations define the prognosis and molecular profiles of patients with gliomas: a meta-analysis." PloS one 8.7 (2013): e68782. https://doi.org/10.1371/journal.pone.0068782
This study, through a meta-analysis, found that IDH1/2 mutations in gliomas are significantly associated with longer overall survival and progression-free survival, and are important potential prognostic biomarkers.
4. Chen, Xiang-Lei, and Shan-Shan Zeng. "Acute myeloid leukemia with NPM1, IDH2, and SETD2 mutations mimicking acute promyelocytic leukemia: A case report and literature review." Medicine 103.42 (2024): e40222. https://doi.org/10.1097/MD.0000000000040222
This case report presents a case of acute myeloid leukemia with mutations in NPM1, IDH2 and SETD2. Its morphology and immunophenotype are similar to acute promyelocytic leukemia, but due to the absence of the PML::RARA fusion gene, it was diagnosed as APL-like AML with NPM1 mutation.
5. Rodriguez, Erika F., et al. "IDH1 and IDH2 mutations in lung adenocarcinomas: evidences of subclonal evolution." Cancer medicine 9.12 (2020): 4386-4394. https://doi.org/10.1002/cam4.3058
This study identified rare (0.5%) IDH1/2 mutations in non-small cell lung cancer, which are commonly found in high-grade adenocarcinoma of elderly smokers and coexist with other driver genes such as KRAS. These mutations may act as branching drivers to promote tumor subclonal evolution.
Creative Biolabs: IDH2 Antibodies for Research
Creative Biolabs specializes in the production of high-quality IDH2 antibodies for research and industrial applications. Our portfolio includes monoclonal antibodies tailored for ELISA, Flow Cytometry, Western blot, immunohistochemistry, and other diagnostic methodologies.
- Custom IDH2 Antibody Development: Tailor-made solutions to meet specific research requirements.
- Bulk Production: Large-scale antibody manufacturing for industry partners.
- Technical Support: Expert consultation for protocol optimization and troubleshooting.
- Aliquoting Services: Conveniently sized aliquots for long-term storage and consistent experimental outcomes.
For more details on our IDH2 antibodies, custom preparations, or technical support, contact us at email.
Reference
- Cupp, J. R., and L. McAlister-Henn. "NAD (+)-dependent isocitrate dehydrogenase. Cloning, nucleotide sequence, and disruption of the IDH2 gene from Saccharomyces cerevisiae." Journal of Biological Chemistry 266.33 (1991): 22199-22205. https://doi.org/10.1016/S0021-9258(18)54554-3
Anti-IDH2 antibodies
 Loading...
Loading...
Hot products 
-
Mouse Anti-ABCA3 Recombinant Antibody (V2-178911) (CBMAB-A0145-YC)

-
Rabbit Anti-ATF4 Recombinant Antibody (D4B8) (CBMAB-A3872-YC)

-
Human Anti-SARS-CoV-2 S1 Monoclonal Antibody (CBFYR-0120) (CBMAB-R0120-FY)

-
Mouse Anti-ARSA Recombinant Antibody (CBYC-A799) (CBMAB-A3679-YC)

-
Mouse Anti-BRCA2 Recombinant Antibody (CBYY-1728) (CBMAB-2077-YY)

-
Mouse Anti-Acetyl SMC3 (K105/K106) Recombinant Antibody (V2-634053) (CBMAB-AP052LY)

-
Mouse Anti-COL1A2 Recombinant Antibody (CF108) (V2LY-1206-LY626)

-
Mouse Anti-ADAM29 Recombinant Antibody (V2-179787) (CBMAB-A1149-YC)

-
Mouse Anti-ACVR1C Recombinant Antibody (V2-179685) (CBMAB-A1041-YC)

-
Mouse Anti-ATP1B3 Recombinant Antibody (1E9) (CBMAB-A4021-YC)

-
Mouse Anti-BBS2 Recombinant Antibody (CBYY-0253) (CBMAB-0254-YY)

-
Mouse Anti-8-oxoguanine Recombinant Antibody (V2-7719) (CBMAB-1898CQ)

-
Rabbit Anti-ABL1 (Phosphorylated Y245) Recombinant Antibody (V2-505716) (PTM-CBMAB-0465LY)

-
Mouse Anti-ATP1B1 Recombinant Antibody (E4) (CBMAB-0463-LY)

-
Rabbit Anti-BAD (Phospho-Ser136) Recombinant Antibody (CAP219) (CBMAB-AP536LY)

-
Mouse Anti-AKR1B1 Antibody (V2-2449) (CBMAB-1001CQ)

-
Mouse Anti-CD33 Recombinant Antibody (P67.6) (CBMAB-C10189-LY)

-
Mouse Anti-AGO2 Recombinant Antibody (V2-634169) (CBMAB-AP203LY)

-
Rat Anti-(1-5)-α-L-Arabinan Recombinant Antibody (V2-501861) (CBMAB-XB0003-YC)

-
Mouse Anti-ALDOA Recombinant Antibody (A2) (CBMAB-A2316-YC)

- AActivation
- AGAgonist
- APApoptosis
- BBlocking
- BABioassay
- BIBioimaging
- CImmunohistochemistry-Frozen Sections
- CIChromatin Immunoprecipitation
- CTCytotoxicity
- CSCostimulation
- DDepletion
- DBDot Blot
- EELISA
- ECELISA(Cap)
- EDELISA(Det)
- ESELISpot
- EMElectron Microscopy
- FFlow Cytometry
- FNFunction Assay
- GSGel Supershift
- IInhibition
- IAEnzyme Immunoassay
- ICImmunocytochemistry
- IDImmunodiffusion
- IEImmunoelectrophoresis
- IFImmunofluorescence
- IGImmunochromatography
- IHImmunohistochemistry
- IMImmunomicroscopy
- IOImmunoassay
- IPImmunoprecipitation
- ISIntracellular Staining for Flow Cytometry
- LALuminex Assay
- LFLateral Flow Immunoassay
- MMicroarray
- MCMass Cytometry/CyTOF
- MDMeDIP
- MSElectrophoretic Mobility Shift Assay
- NNeutralization
- PImmunohistologyp-Paraffin Sections
- PAPeptide Array
- PEPeptide ELISA
- PLProximity Ligation Assay
- RRadioimmunoassay
- SStimulation
- SESandwich ELISA
- SHIn situ hybridization
- TCTissue Culture
- WBWestern Blot






