SLIRP Antibodies
Background
The SLIRP gene encodes a RNA-binding protein, which is mainly located in the cell nucleus and mitochondria. This protein specifically binds to the stem-loop structure of the nuclear receptor co-activator SRA, participating in the regulation of transcriptional activities mediated by nuclear hormone receptors, and influencing the assembly and function of the respiratory chain complex within mitochondria. Studies have found that SLIRP plays an important role in maintaining the balance of cellular energy metabolism, oxidative stress response, and embryonic development. Its abnormal expression is associated with various metabolic diseases and cancer progression. As a key node in the epigenetic and metabolic regulation network, the mechanism of SLIRP's action continuously provides new perspectives for the study of cell signal transduction.
Structure of SLIRP
SLIRP is a relatively small RNA-binding protein with a molecular weight of approximately 18.5 kDa. This molecular weight varies slightly among different species or different isoforms of the same species, mainly due to the minor changes in amino acid sequence caused by its alternative splicing.
| Species | Human (Heteromer 1) | Mouse (main isomer) |
| Molecular Weight (kDa) | About 18.5 | About 18.3 |
| Primary Structural Differences | With nuclear localization signal and mitochondrial targeting sequence | The sequence is highly conserved and the functional domains are similar. |
The SLIRP protein consists of approximately 160 amino acids and forms its functional core through its RNA recognition motif (RRM). The spatial conformation of this protein enables it to specifically recognize and bind to the specific stem-loop structure of the nuclear receptor co-activator SRA RNA. The α helices and β sheets in its structure jointly maintain a binding interface, which is crucial for its function in regulating gene transcription and participating in mitochondrial RNA metabolism.
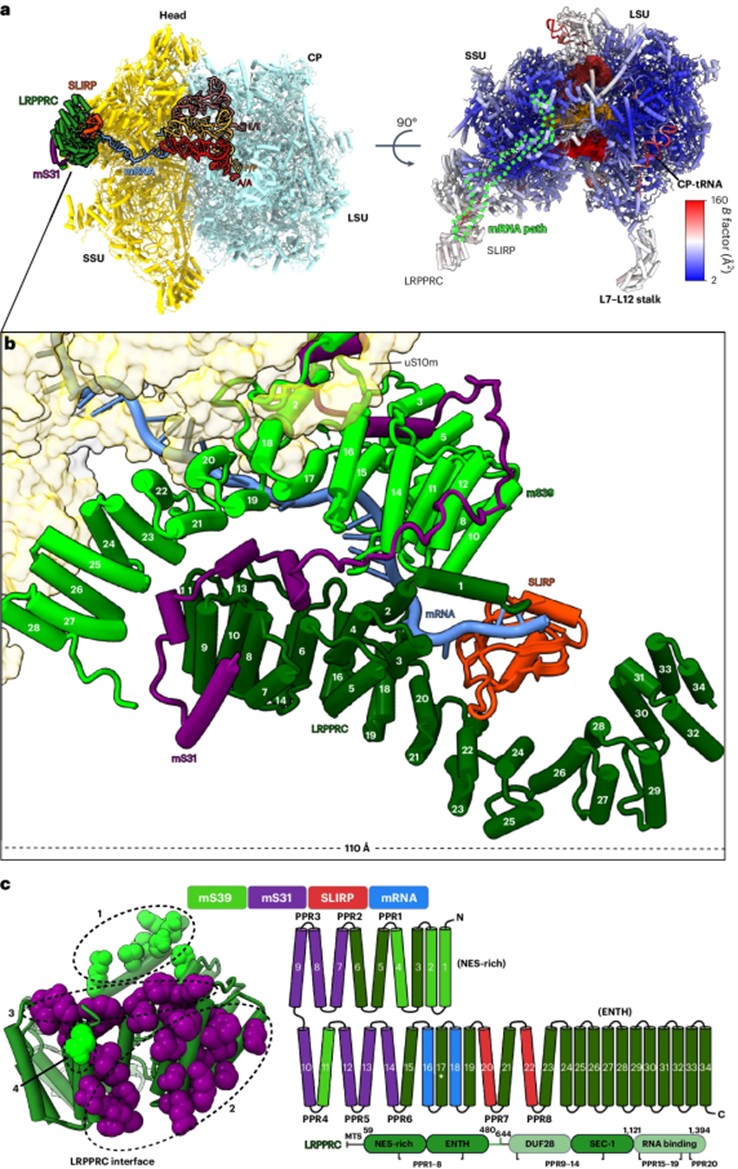 Fig. 1 Structure of mitoribosome with LRPPRC–SLIRP bound to mRNA.1
Fig. 1 Structure of mitoribosome with LRPPRC–SLIRP bound to mRNA.1
Key structural properties of SLIRP:
- Contains a classic RNA recognition motif (RRM)
- Core structure specificity of recognition and combination of SRA RNA stem loop structure
- Have approved a signal and mitochondria double location information of target sequence
Functions of SLIRP
The main function of the SLIRP gene is to act as a dual regulator for transcription and mitochondrial function. The specific physiological processes it is involved in are as follows:
| Function | Description |
| Transcriptional Regulation | By specifically binding to the stem-loop structure of SRA RNA, it regulates the transcriptional activity of genes mediated by nuclear receptors (such as estrogen receptor, thyroid hormone receptor). |
| Mitochondrial Function | Located within the mitochondria, it participates in the metabolism and stability of mitochondrial RNA, and affects the assembly of respiratory chain complexes as well as the efficiency of cellular energy metabolism. |
| Metabolic Balance | As a co-regulatory factor, the signaling pathways involved in processes such as glucose metabolism and lipid metabolism have abnormal expressions, which are associated with metabolic diseases. |
| Developmental Regulation | Expressed during embryonic development, it plays a crucial role in normal tissue differentiation and the overall developmental process. |
| Stress Response | It involves the adaptive responses of cells to environmental stresses such as oxidative stress, and affects the survival and function of cells. |
The mechanism of SLIRP highlights its pivotal role as a key node linking nuclear gene expression with mitochondrial energy metabolism. This cross-organelle coordination function is of utmost importance for maintaining cellular homeostasis.
Applications of SLIRP and SLIRP Antibody in Literature
1. Singh, Vivek, et al. "Structural basis of LRPPRC–SLIRP-dependent translation by the mitoribosome." Nature structural & molecular biology 31.12 (2024): 1838-1847. https://doi.org/10.1038/s41594-024-01365-9
The mitochondrial factor LRPPRC forms a complex with SLIRP. Through structural analysis, it was found that this complex can bind to mRNA and assist in its delivery to the mitochondrial ribosomes. Studies have shown that this complex can regulate the translation efficiency of different mRNAs, especially affecting the synthesis of cytochrome c oxidase subunits, and is an important regulator of mitochondrial gene expression.
2. Colley, Shane M., et al. "Loss of the nuclear receptor corepressor SLIRP compromises male fertility." PLoS One 8.8 (2013): e70700. https://doi.org/10.1371/journal.pone.0070700
This study reveals that the SLIRP protein not only inhibits the activity of nuclear receptors, but also is abnormally distributed in the testes. In genetically knockout mice, the absence of SLIRP leads to a decline in male fertility, impaired sperm motility and mitochondrial structure, thereby causing asthenospermia.
3. De Silva, Dinuka, et al. "Interaction between androgen receptor and coregulator SLIRP is regulated by Ack1 tyrosine kinase and androgen." Scientific Reports 9.1 (2019): 18637. https://doi.org/10.1038/s41598-019-55057-2
This study reveals that SLIRP, as a co-regulator of the androgen receptor (AR), binds to target genes in the absence of androgens. The Ack1 kinase and androgen signaling can disrupt the binding of AR-SLIRP, affecting the transcription of related genes, suggesting its role in castration-resistant prostate cancer.
4. Lagouge, Marie, et al. "SLIRP regulates the rate of mitochondrial protein synthesis and protects LRPPRC from degradation." PLoS genetics 11.8 (2015): e1005423. https://doi.org/10.1371/journal.pgen.1005423
This study reveals that SLIRP mainly stabilizes the LRPPRC protein in the LRPPRC-SLIRP complex and is responsible for mediating the binding of mitochondrial mRNA to ribosomes, thereby regulating translation efficiency. The study found that mitochondrial mRNA exhibits significant redundancy under normal physiological conditions.
5. Baleva, Mariia V., et al. "Mitochondrial Protein SLIRP Affects Biosynthesis of Cytochrome c Oxidase Subunits in HEK293T Cells." International Journal of Molecular Sciences 25.1 (2023): 93. https://doi.org/10.3390/ijms25010093
This study reveals that the SLIRP protein specifically binds to the small subunit of mitochondrial ribosomes and regulates translation initiation. Its absence interferes with mitochondrial translation, leading to dysfunction of complexes I and IV, but does not affect complexes III and V.
Creative Biolabs: SLIRP Antibodies for Research
Creative Biolabs specializes in the production of high-quality SLIRP antibodies for research and industrial applications. Our portfolio includes monoclonal antibodies tailored for ELISA, Flow Cytometry, Western blot, immunohistochemistry, and other diagnostic methodologies.
- Custom SLIRP Antibody Development: Tailor-made solutions to meet specific research requirements.
- Bulk Production: Large-scale antibody manufacturing for industry partners.
- Technical Support: Expert consultation for protocol optimization and troubleshooting.
- Aliquoting Services: Conveniently sized aliquots for long-term storage and consistent experimental outcomes.
For more details on our SLIRP antibodies, custom preparations, or technical support, contact us at email.
Reference
- Singh, Vivek, et al. "Structural basis of LRPPRC–SLIRP-dependent translation by the mitoribosome." Nature structural & molecular biology 31.12 (2024): 1838-1847. Distributed under Open Access license CC BY 4.0, without modification.https://doi.org/10.1038/s41594-024-01365-9
Anti-SLIRP antibodies
Products List
 Loading...
Loading...
Hot products 
-
Mouse Anti-dsRNA Recombinant Antibody (2) (CBMAB-D1807-YC)

-
Mouse Anti-BZLF1 Recombinant Antibody (BZ.1) (CBMAB-AP705LY)

-
Mouse Anti-AQP2 Recombinant Antibody (G-3) (CBMAB-A3359-YC)

-
Mouse Anti-ARSA Recombinant Antibody (CBYC-A799) (CBMAB-A3679-YC)

-
Mouse Anti-ABIN2 Recombinant Antibody (V2-179106) (CBMAB-A0349-YC)

-
Mouse Anti-FAS2 Monoclonal Antibody (1D4) (CBMAB-0071-CN)

-
Mouse Anti-CD33 Recombinant Antibody (6C5/2) (CBMAB-C8126-LY)

-
Mouse Anti-CARD11 Recombinant Antibody (CBFYC-0811) (CBMAB-C0866-FY)

-
Mouse Anti-CASP7 Recombinant Antibody (10-01-62) (CBMAB-C2005-LY)

-
Rabbit Anti-Acetyl-Histone H3 (Lys36) Recombinant Antibody (V2-623395) (CBMAB-CP0994-LY)

-
Mouse Anti-AAV-5 Recombinant Antibody (V2-503417) (CBMAB-V208-1369-FY)

-
Mouse Anti-BACE1 Recombinant Antibody (61-3E7) (CBMAB-1183-CN)

-
Mouse Anti-BHMT Recombinant Antibody (CBYY-0547) (CBMAB-0550-YY)

-
Mouse Anti-ADAM12 Recombinant Antibody (V2-179752) (CBMAB-A1114-YC)

-
Mouse Anti-FLI1 Recombinant Antibody (CBXF-0733) (CBMAB-F0435-CQ)

-
Mouse Anti-APOH Recombinant Antibody (4D9A4) (CBMAB-A3249-YC)

-
Mouse Anti-AFM Recombinant Antibody (V2-634159) (CBMAB-AP185LY)

-
Rat Anti-ADGRE4 Recombinant Antibody (V2-160163) (CBMAB-F0011-CQ)

-
Mouse Anti-AKR1B1 Antibody (V2-2449) (CBMAB-1001CQ)

-
Mouse Anti-ARHGAP5 Recombinant Antibody (54/P190-B) (CBMAB-P0070-YC)

- AActivation
- AGAgonist
- APApoptosis
- BBlocking
- BABioassay
- BIBioimaging
- CImmunohistochemistry-Frozen Sections
- CIChromatin Immunoprecipitation
- CTCytotoxicity
- CSCostimulation
- DDepletion
- DBDot Blot
- EELISA
- ECELISA(Cap)
- EDELISA(Det)
- ESELISpot
- EMElectron Microscopy
- FFlow Cytometry
- FNFunction Assay
- GSGel Supershift
- IInhibition
- IAEnzyme Immunoassay
- ICImmunocytochemistry
- IDImmunodiffusion
- IEImmunoelectrophoresis
- IFImmunofluorescence
- IGImmunochromatography
- IHImmunohistochemistry
- IMImmunomicroscopy
- IOImmunoassay
- IPImmunoprecipitation
- ISIntracellular Staining for Flow Cytometry
- LALuminex Assay
- LFLateral Flow Immunoassay
- MMicroarray
- MCMass Cytometry/CyTOF
- MDMeDIP
- MSElectrophoretic Mobility Shift Assay
- NNeutralization
- PImmunohistologyp-Paraffin Sections
- PAPeptide Array
- PEPeptide ELISA
- PLProximity Ligation Assay
- RRadioimmunoassay
- SStimulation
- SESandwich ELISA
- SHIn situ hybridization
- TCTissue Culture
- WBWestern Blot






