EPAS1 Antibodies
Background
The EPAS1 gene encodes hypoxia-inducible factor 2α (HIF-2α), which is a type of transcription factor that plays a central regulatory role in low-oxygen environments. This protein is mainly expressed in vascular endothelial cells, cardiomyocytes, and certain neuroendocrine cells, and it activates the expression of downstream genes by sensing changes in oxygen concentration, thereby regulating physiological processes such as the body's oxygen balance adaptation, angiogenesis, and erythropoiesis. This gene was first identified in 1997, and the elucidation of its function and structure not only deepened our understanding of the biological response to hypoxia but also provided an important foundation for the research on target points for cardiovascular diseases and tumor treatment.
Structure of EPAS1
The molecular weight of the EPAS1 protein (the α subunit of hypoxia-inducible factor) is approximately 100 kDa. The number of amino acid residues varies among different species, resulting in a slight fluctuation in molecular weight.
| Species | Human | Mouse | Zang people (adaptive variations) | Bovine |
|---|---|---|---|---|
| Molecular Weight (kDa) | 100 | 98 | 100 | 99 |
| Primary Structural Differences | Containing a PAS domain and a DNA binding domain, with an oxygen-sensitive degradation region | Homologous structure highly conservative, some different amino acids | Mutations at key sites (such as Pro531, etc.) affect the intensity of the hypoxia response. | Similar to the structure of humans and having similar functions |
This protein is composed of approximately 870 amino acid residues and exhibits a modular structure. Its N-terminal region contains a helix-loop-helix (HLH) and two tandem PAS (Per-ARNT-Sim) domains, which are responsible for dimerization with the ARNT protein and recognition of hypoxia response elements on DNA; the oxygen-dependent degradation domain (ODD) at the C-terminal is ubiquitinated and degraded by proline hydroxylase under normoxic conditions. The protein's internal hydrophobic core encapsulates the interface that binds to cofactors (such as HRE sequences), and key sites such as specific residues within the PAS-B domain (for example, the common D3E variation among Zang people) can alter protein stability or transcriptional activity, thereby affecting the organism's ability to adapt to low-oxygen environments. Its activity is directly regulated by cellular oxygen concentration and constitutes the core of oxygen homeostasis regulation.
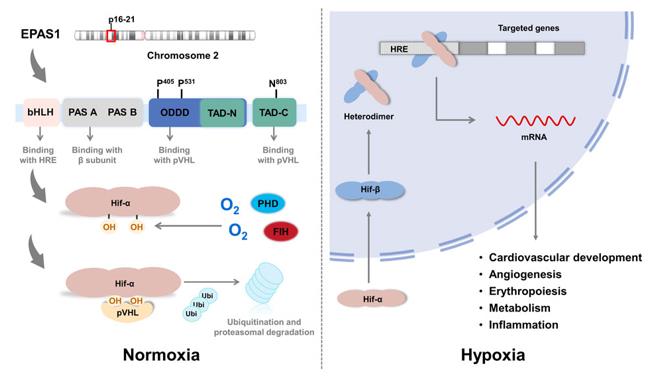 Fig. 1 The structure of EPAS1.1
Fig. 1 The structure of EPAS1.1
Key structural properties of EPAS1:
- Modular protein domains (including helix-ring-helix and PAS functional domains)
- Stability regulation mediated by the oxygen-sensitive degradation domain (ODD)
- Key amino acid sites (such as Tibetan-specific variant sites) affect the efficiency of hypoxia response
Functions of EPAS1
The core function of the EPAS1 gene (also known as HIF-2α) is to act as the primary regulatory factor in hypoxic conditions, coordinating the body's adaptation and response to oxygen deprivation. Its main physiological functions are as follows:
| Function | Description |
|---|---|
| Oxygen deprivation response regulation | Under hypoxic conditions, it is stably expressed and translocated to the nucleus, forming a dimer with ARNT protein. It binds to the hypoxia response element (HRE) of downstream target genes and initiates transcription. |
| Red Blood Cell Generation Regulation | Promotes the expression of genes such as erythropoietin (EPO), stimulates red blood cell production, and enhances the oxygen-carrying capacity of the blood. |
| Induction of Angiogenesis | Upregulate genes such as vascular endothelial growth factor (VEGF) to promote the formation of new blood vessels and improve tissue oxygen supply. |
| Energy Metabolism Reprogramming | Regulates genes related to glycolysis, helping cells switch to anaerobic metabolism when oxygen is scarce to maintain energy supply. |
| Key Role of Altitude Adaptation | Compared with other members of the HIF family (such as HIF-1α), EPAS1 has a more specific role in regulating specific physiological processes (such as pulmonary vascular responses, catecholamine metabolism, etc.). |
Its expression and activity are precisely regulated by oxygen concentration, prolyl hydroxylase, and the ubiquitination degradation pathway.
Applications of EPAS1 and EPAS1 Antibody in Literature
1. Pirri, Daniela, et al. "EPAS1 attenuates atherosclerosis initiation at disturbed flow sites through endothelial fatty acid uptake." Circulation research 135.8 (2024): 822-837. https://doi.org/10.1161/CIRCRESAHA.123.324054
Research has revealed the protective effect of the transcription factor EPAS1 in atherosclerosis: the expression of EPAS1 increases in areas with blood flow disorders, and obesity inhibits its expression through the triglyceride-PHD2 pathway, leading to a decline in the proliferation and repair ability of endothelial cells, thereby accelerating plaque formation. Targeting this pathway may provide a new strategy for obesity-related vascular lesions.
2. Kyriakopoulou, Eirini, et al. "EPAS1 induction drives myocardial degeneration in desmoplakin-cardiomyopathy." Iscience 28.3 (2025). https://doi.org/10.1016/j.isci.2025.111895
Research has revealed that EPAS1 is a key regulatory factor driving mitochondrial dysfunction, hypoxia stress and myocardial degeneration in cardiomyocytes in arrhythmic cardiomyopathy (ACM). This discovery provides new molecular mechanisms and potential therapeutic targets for ACM related to gene mutations such as DSP.
3. Wang, Na, et al. "Updated perspective of EPAS1 and the role in pulmonary hypertension." Frontiers in cell and developmental biology 11 (2023): 1125723. https://doi.org/10.3389/fcell.2023.1125723
Research reveals that the core pathology of pulmonary arterial hypertension (PH) is hypoxia-driven vascular remodeling. This article reviews the molecular structure and function of EPAS1 (HIF-2α) and its regulatory network in cardiovascular diseases such as PH, providing a theoretical basis for the development of new targets for vascular remodeling.
4. Li, Qinyu, et al. "Atractylenolide I inhibits angiogenesis and reverses sunitinib resistance in clear cell renal cell carcinoma through ATP6V0D2-mediated autophagic degradation of EPAS1/HIF2α."Autophagy21.3 (2025): 619-638. https://doi.org/10.1080/15548627.2024.2421699
Studies have revealed that Atletalide I (ATL-I) inhibits angiogenesis and tumor growth in clear cell renal cell carcinoma (ccRCC) by upregulating ATP6V0D2, promoting autophagic degradation of EPAS1 (HIF-2α), and suppressing VEGFA production, and can reverse sunitinib resistance.
5. Liu, Lili, et al. "EPAS1 amplifies asthma pathogenesis through JAK2/STAT3-mediated ferroptosis and inflammation." Biomolecules and Biomedicine 25.9 (2025): 2092. https://doi.org/10.17305/bb.2025.11334
Research has found that in asthma, the JAK2/STAT3 signaling axis drives ferroptosis and inflammatory responses by up-regulating EPAS1 expression. EPAS1 has been identified as a key hub gene related to immune cell infiltration and can serve as a potential therapeutic target and novel biomarker for asthma.
Creative Biolabs: EPAS1 Antibodies for Research
Creative Biolabs specializes in the production of high-quality EPAS1 antibodies for research and industrial applications. Our portfolio includes monoclonal antibodies tailored for ELISA, Flow Cytometry, Western blot, immunohistochemistry, and other diagnostic methodologies.
- Custom EPAS1 Antibody Development: Tailor-made solutions to meet specific research requirements.
- Bulk Production: Large-scale antibody manufacturing for industry partners.
- Technical Support: Expert consultation for protocol optimization and troubleshooting.
- Aliquoting Services: Conveniently sized aliquots for long-term storage and consistent experimental outcomes.
For more details on our EPAS1 antibodies, custom preparations, or technical support, contact us at email.
Reference
- Wang, Na, et al. "Updated perspective of EPAS1 and the role in pulmonary hypertension." Frontiers in cell and developmental biology 11 (2023): 1125723. https://doi.org/10.3389/fcell.2023.1125723
Anti-EPAS1 antibodies
 Loading...
Loading...
Hot products 
-
Mouse Anti-CD59 Recombinant Antibody (CBXC-2097) (CBMAB-C4421-CQ)

-
Mouse Anti-C5b-9 Recombinant Antibody (aE11) (CBMAB-AO138LY)

-
Mouse Anti-CARTPT Recombinant Antibody (113612) (CBMAB-C2450-LY)

-
Mouse Anti-ADV Recombinant Antibody (V2-503423) (CBMAB-V208-1364-FY)

-
Mouse Anti-CHRNA9 Recombinant Antibody (8E4) (CBMAB-C9161-LY)

-
Mouse Anti-CSPG4 Recombinant Antibody (CBFYM-1050) (CBMAB-M1203-FY)

-
Mouse Anti-CECR2 Recombinant Antibody (CBWJC-2465) (CBMAB-C3533WJ)

-
Mouse Anti-G6PD Recombinant Antibody (13B331) (CBMAB-G1553-LY)

-
Mouse Anti-CD33 Recombinant Antibody (6C5/2) (CBMAB-C8126-LY)

-
Mouse Anti-ALPL Antibody (B4-78) (CBMAB-1009CQ)

-
Mouse Anti-BCL2L1 Recombinant Antibody (H5) (CBMAB-1025CQ)

-
Mouse Anti-ACLY Recombinant Antibody (V2-179314) (CBMAB-A0610-YC)

-
Mouse Anti-BRCA2 Recombinant Antibody (CBYY-0790) (CBMAB-0793-YY)

-
Mouse Anti-AKR1B1 Antibody (V2-2449) (CBMAB-1001CQ)

-
Mouse Anti-CD19 Recombinant Antibody (CBXC-1224) (CBMAB-C1491-CQ)

-
Mouse Anti-ANXA7 Recombinant Antibody (A-1) (CBMAB-A2941-YC)

-
Mouse Anti-AGK Recombinant Antibody (V2-258056) (CBMAB-M0989-FY)

-
Mouse Anti-FLI1 Recombinant Antibody (CBXF-0733) (CBMAB-F0435-CQ)

-
Mouse Anti-APOH Recombinant Antibody (4D9A4) (CBMAB-A3249-YC)

-
Mouse Anti-ALOX5 Recombinant Antibody (33) (CBMAB-1890CQ)

- AActivation
- AGAgonist
- APApoptosis
- BBlocking
- BABioassay
- BIBioimaging
- CImmunohistochemistry-Frozen Sections
- CIChromatin Immunoprecipitation
- CTCytotoxicity
- CSCostimulation
- DDepletion
- DBDot Blot
- EELISA
- ECELISA(Cap)
- EDELISA(Det)
- ESELISpot
- EMElectron Microscopy
- FFlow Cytometry
- FNFunction Assay
- GSGel Supershift
- IInhibition
- IAEnzyme Immunoassay
- ICImmunocytochemistry
- IDImmunodiffusion
- IEImmunoelectrophoresis
- IFImmunofluorescence
- IGImmunochromatography
- IHImmunohistochemistry
- IMImmunomicroscopy
- IOImmunoassay
- IPImmunoprecipitation
- ISIntracellular Staining for Flow Cytometry
- LALuminex Assay
- LFLateral Flow Immunoassay
- MMicroarray
- MCMass Cytometry/CyTOF
- MDMeDIP
- MSElectrophoretic Mobility Shift Assay
- NNeutralization
- PImmunohistologyp-Paraffin Sections
- PAPeptide Array
- PEPeptide ELISA
- PLProximity Ligation Assay
- RRadioimmunoassay
- SStimulation
- SESandwich ELISA
- SHIn situ hybridization
- TCTissue Culture
- WBWestern Blot






